Multi-Objective Feature Selection Part 4: Making Better Machine Learning Models
 sgenzer
Administrator, Moderator, Employee, RapidMiner Certified Analyst, Community Manager, Member, University Professor, PM Moderator Posts: 2,959
sgenzer
Administrator, Moderator, Employee, RapidMiner Certified Analyst, Community Manager, Member, University Professor, PM Moderator Posts: 2,959 
Feature Selection for Unsupervised Learning
In part 1 of this series, we established that feature selection is a computationally hard problem. We then saw that evolutionary algorithms can tackle this problem in part 2. Finally, we discussed and that multi-objective optimization delivers additional insights into your data and machine learning model.
There’s one thing we haven’t discussed yet which is multi-objective feature selection. It can also be used for unsupervised learning. This means that you can now also identify the best feature spaces in which to find your clusters. Let’s discuss the problem in more detail and see how we can now solve it.
Unsupervised Feature Selection and the Density Problem
We will focus on clustering problems for this post. Everything below is valid for most other unsupervised learning techniques as well.
Ok, let’s discuss k-means clustering for now. The idea of the algorithm is to identify centroids for a given number of clusters. Those centroids are the average of all features of the data points of their respective cluster. We then assign all data points to those centroids which – after some points have been re-assigned – will be recalculated. This procedure stops after either a maximal number of iterations or after the clusters stay stable and no point has been re-assigned.
So how can we measure how well the algorithm segmented our data? There is a common technique for this, namely the Davis-Bouldin index (or short: DB index). It can be calculated using the following formula:

with n as the number of clusters, ci as centroid of cluster i, σi as the average distance of all points of cluster ito their centroid, and d(ci,cj) as the distance between the centroids of clusters i and j.
As we can see, the result of a clustering is better if points within a cluster are close to each other. That of course is exactly what we wanted, but this also means that the DB index prefers clustering results where the clusters have a higher density.
But this preference for higher densities in the clusters is posing a problem for feature selection. If we do feature selection, we reduce the number of features. Less features also means higher densities in the remaining dimensions. This makes intuitive sense, because we map the data points from the higher-dimensional space into a smaller number of dimensions, bringing the points closer to each other.
Traditional feature selection cannot be used for clustering. Imagine that we have one feature as part of our data that is pure noise, but completely random. Say we flip a coin and use 0 for head and 1 for tails. We want to use k-means clustering with k=2 to find two clusters in our data and then decide to use forward selection to find the best feature set for this task. It first tries to find the best clusters using only one feature. Of course, our random feature described above will win this race: it will produce two clusters with infinite density! One cluster will have all the points at 0 and the other one all the points at 1, but keep in mind that this was a complete random feature. As a result, clustering only using this dimension would be completely meaningless for the original problem and data space.
Even in cases where we don’t have a pseudo-categorical random feature, we will always pick only one feature. Namely the one which delivers the k densest clusters. Adding more features would only reduce the density again so we will not go there.
The Solution: Multi-objective Feature Selection
Can multi-objective optimization help with this problem? Let’s do some experiments and see how this works. The data set below has four clusters in two dimensions: att1 and att2. There are also four additional random features called random1, random2 etc. The random features use a gaussian distribution of values around 0. Please note: the data is normalized so all data columns have mean 0 and standard deviation 1. This process is called standardization.
Links to each of these processes can be found at the bottom of this post if you want to follow along. Here is how the data set looks if you only look at the dimensions att1 and att2:

It is easy to see the four natural clusters here. However, things change a bit if you plot the same data with the same point colors. Below we use the dimensions random1 and random2 instead:

As expected, you can no longer see any clusters, and the data points are randomly scattered around the center. Before we even start with feature selection, let’s just run a k-means clustering with k=4 on the full data set consisting of all 6 features.
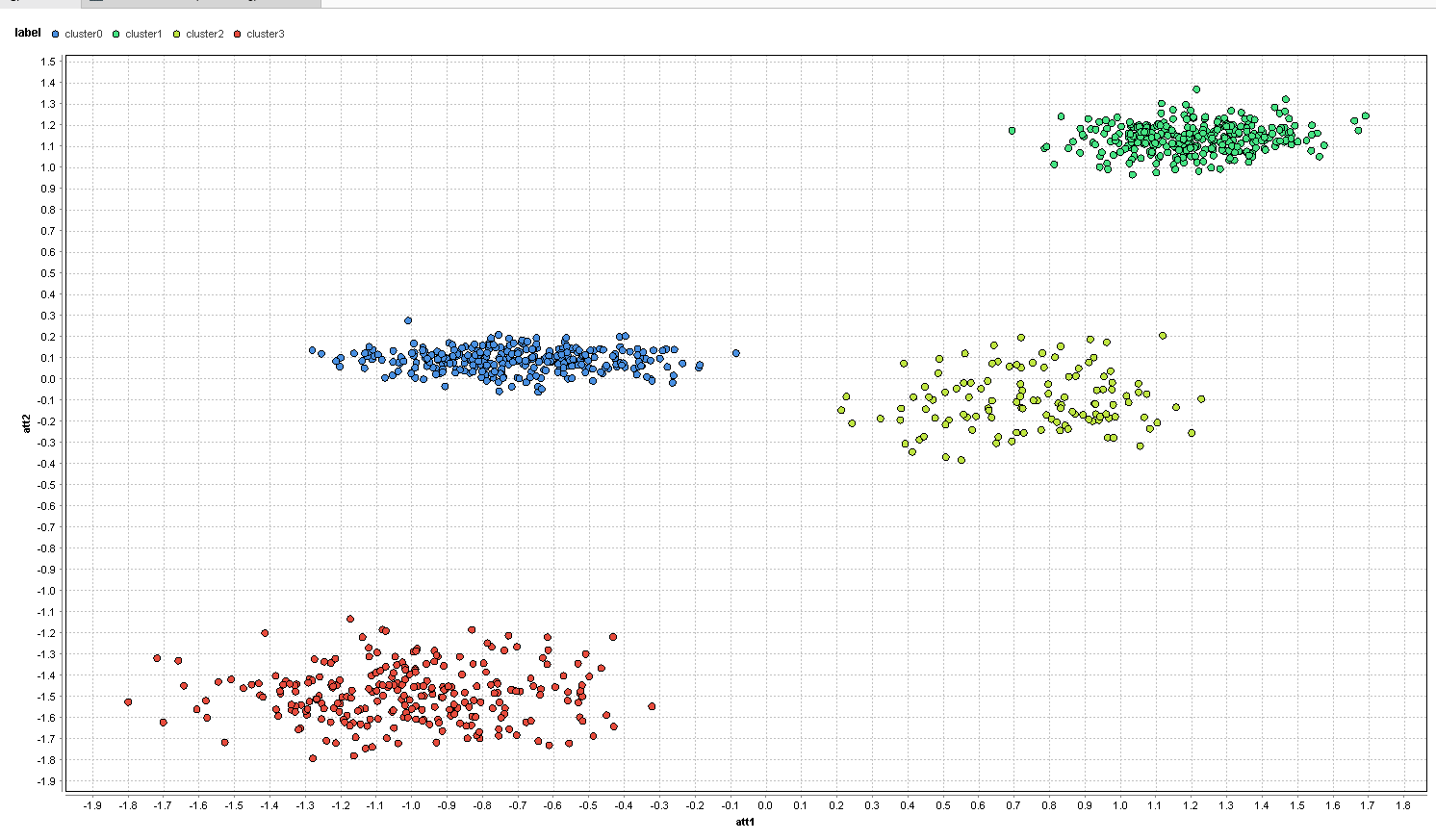
The clusters now represent the clusters found by k-means and they are not completely horrible. Still, the additional random attributes managed to throw off our k-means. On the left, you have the red and blue clusters completely mixed up and some of the red and blue points are even in the right clusters. Same is true for the yellow and green clusters on the right.
We can see that the noise attributes can render the clustering completely meaningless. The desired clusters have not been found. Since traditional feature selection is not helpful, let’s see how well a multi-objective approach will do.
We use the same basic setup like in the previous post (LINK). First, we retrieve the data set and then normalize the data as described above. We then use an evolutionary feature selection scheme where we use “non-dominated sorting” as the selection scheme.

The differences are a bit bigger inside of the feature selection operator:

Instead of using a 10-fold cross validation, we can create the clustering using k=4. The first performance operator takes the clustering result and calculates the DB-Index. The second operator calculates the number of features used for the current individual. We add this number as the second objective for our optimization.
We can execute this process now. Below is the resulting Pareto front after 30 generations:
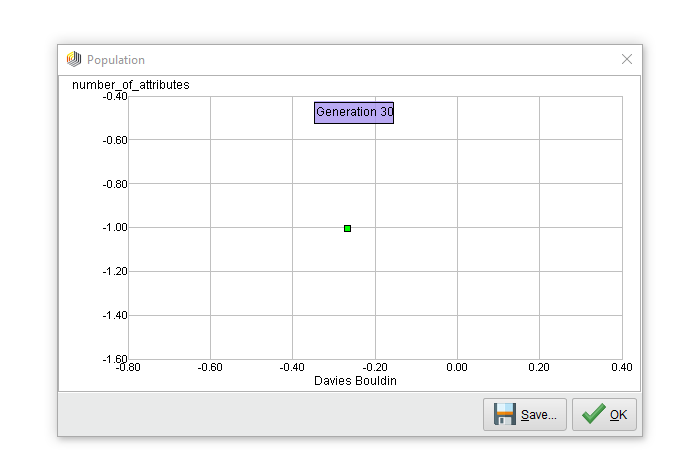
Only one point on the Pareto front? Of course! Any feature selection scheme which tries to minimize the number of features and optimize the density of clusters will always end up with only one single feature. The Pareto front collapsed into this single point after 12 generations and stayed that way for the remaining run.
The selected feature was att2 which is not a bad choice at all. In fact, here is the histogram showing the distributions for the four found clusters in that dimension:

The red and green clusters are pretty good. However, there is a lot of overlap between the blue and the yellow clusters. Check out the first image in this post again and project the four clusters to only att2 and it will be the same result.
We can concur that we need both attributes att2 and att1 to get to the desired clusters, but we ended up with only one feature instead. It seems that multi-objective optimization alone does not help here. The reason is that the two goals are not conflicting so there will not be a Pareto front showing all trade-offs. There are simply no trade-offs to show.
Here is an idea: What if we change the optimization direction for the number of features? If minimizing the number of features is not conflicting, why are we not introducing a conflict to maximize the number instead?
This might sound crazy at first, but it makes sense if we think about it. The whole point of clustering is to describe your data. It would be good to stay as close to the original data space as possible and only omit what is truly garbage. If we omit too much, then our description would not be useful anymore. It is like omitting too many details when you describe an object. “It has wheels” is not a wrong description of “cars”, but it also applies to many other things like bicycles, motorbikes, and even airplanes. But adding “…and four seats and an engine” makes this a much better description without adding too many unnecessary details. In my PhD thesis, we called this idea “information preservation.”
In fact, you only need to change one thing in the RapidMiner process above. The second performance operator has a parameter creatively called “optimization direction.” When we change this from “minimize” to “maximize” and we are ready to go. Below is the Pareto front we are getting at the end of this optimization run:

That looks much better, doesn’t it? We get five different points on this Pareto front. There is a clear difference for the DB index using only the first two attributes and the remaining solutions using 3, 5, or 6 features. What makes those first two solutions special then? Here is a table with the details about the solutions:

First, we get two times the same solution using only one attribute. We already know that att2 is the winner here having the best DB index. However, the next one is only slightly worse in terms of DB index and adds the “necessary” feature att1 as well. From there on, there is clear drop for the DB-Index when we start adding the random features.
There is no way to make the final choice of a cluster set without inspecting the resulting clusters. But now we can at least see if they make sense and describe our data in a meaningful way. It is often a good idea to start at those areas short before clear drops for the DB-index. This is not dissimilar to the known elbow criterion for selecting the best number k of clusters. We can cleary see the shape of this drop-off in the Pareto front chart above.
Thanks to the information-preserving nature of the approach, it delivers all relevant information. And you get this information in one single optimization run. Feature selection for unsupervised learning has been finally solved.
Conclusion
Even in the age of deep learning, there are still a lot of applications where other model types are superior or preferred. Additionally, in almost all cases those models benefit from reducing the noise in the input data by dropping unnecessary features. Models are less prone to overfitting and become simpler which makes them more robust against small data changes. Also, the simplicity improves understandability of the models a lot.
Evolutionary algorithms are a powerful technique for feature selection. They are not getting stuck in the first local optimum such as forward selection or backward elimination. This leads to more accurate models.
An even better approach is using a multi-objective selection technique. These algorithms deliver the full Pareto front of solutions in the same runtime. We can inspect the trade-off between complexity and accuracy yourself (“is it really worth to add this additional feature?”) and you will get additional insights about the interaction of the features.
Finally, this multi-objective feature selection approach has another advantage. We can use it for unsupervised learning like for clustering techniques. However, we need to change the optimization direction for the number of features. This can be easily seen when maximizing density-based cluster measurements. If we simultaneously also minimize the number of features, we end up with only one single attribute, but if we maximize the number of attributes, it is harder to find dense clusters for more features. This will lead to the best clusters for all sensible numbers of features. We also preserve the original information which is what clustering is all about: find the hidden segments in your original data.
Here we have it: the one and only feature selection solution needed in the future.
RapidMiner Processes
You can download RapidMiner here. Then you can download the processes below to build this machine learning model yourself in RapidMiner. The result will be a .rmp file which can be loaded into RapidMiner via “File” -> “Import Process.” Please note that you need to run the first three processes to create the data sets which will be used by the other processes. Please adapt the storage paths in the parameters of the storage and retrieval operators of all processes according to your setup.
Author: @IngoRM, Founder & President of RapidMiner


Comments
If it is still available, could someone please post a link to the "Part 4" example process? It is referenced in the article and I may be overlooking something obvious, but I don't see it.
Thanks,
Noel
Ingo